HAPLOTYPES, RECESSIVES AND GENETIC CODES EXPLAINED
This page is designed to help explain the current genetic codes which appear after the names of registered Holstein animals on pedigrees, web factsheets and top lists.
Firstly here are a few definitions of common genetic terms, which will be used in the description and explanation of genetic codes. The diagrams below can help visualise these genetic components. A further reading section at the end of this document also alerts you to some useful explanatory material on the subject.
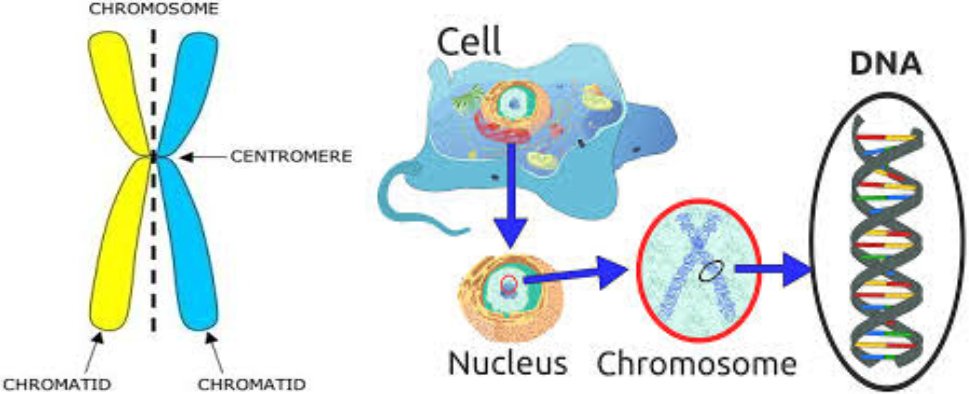
A Gene: is a sequence DNA which codes (i.e. is the instruction manual) for a particular amino acid. These amino acids are the building blocks of proteins. Each animal has two copies of each gene (which can vary - see Allele below).
A Chromosome: is a large unit of DNA, containing many genes. Each chromosome is made up of a pair of chromatids. Cattle have 30 chromosomes found in most cells in the body. In Sperm and Eggs, Chromatids exist as un- paired single entities. Therefore an animal inherits half of each chromosome (and thus one copy of each gene) from each parent.
An Allele: is a form or version of a gene. An animal can have the same allele on both chromosomes (Homozygous) or different alleles (Heterozygous). Genes can have multiple alleles, such as the MCR1 gene responsible for some of the red coat colour variations, however 2 is most common. An animal can only inherit one allele from each parent.
A Recessive allele: is one which only has an effect when an animal inherits this version of the gene from each parent. A dominant allele affects an animal when either one or two versions are inherited. Most harmful alleles are recessive.
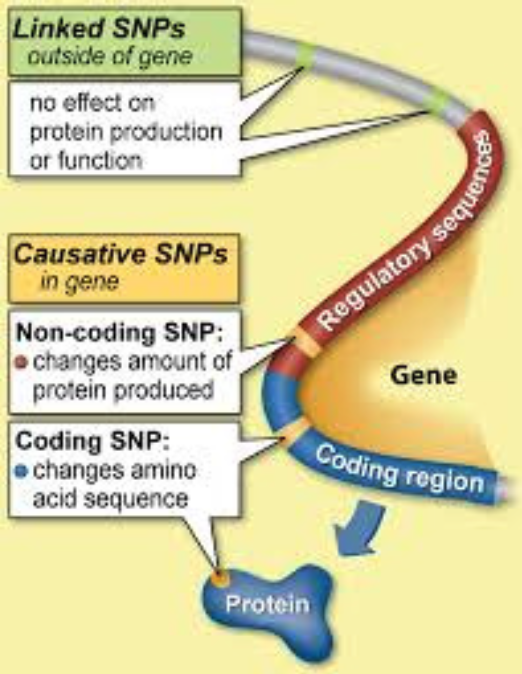
A SNP (Single Nucleotide Polymorphism): is a single unit of DNA which varies between animals within the same breed or species. A SNP may be located within a gene or on a region between genes on a chromosome. SNP are the markers used to genotype animals for genomic selection.
A Haplotype: is a group of SNP or alleles located close to each other on the chromosome, which are usually inherited together.
RECESSIVE DEFECTS
| Gene Name | Description | Gene & Expression Code |
| BLAD | Bovine Leukocyte Adhesion Deficiency (deficiency of a normally occurring protein needed for white blood cells or leukocytes, which are body’s infection fighters) | BLC = tested carrier of BLAD BLF = tested non-carrier of BLAD |
| Mule Foot | Mule-Foot (toes of foot are joined, giving animal a single hoof, instead of cloven ones) | MFC = tested carrier of Mule foot MFF = tested non-carrier of Mule foot |
| DUMPS | Deficiency of Uridine Monophosphate Synthase (one of many enzymes contributing to normal metabolic processes) | DPC = tested carrier of DUMPS DPF = tested non-carrier of DUMPS |
| CVM | Complex Vertebral Malformation (causes still-born calves, abortions, and early embryonic losses) | CVC = tested carrier of CVM CVF = tested non-carrier of CVM |
| Factor X1 | Factor X1 (blood clotting disorder) | XIC = tested carrier of Factor X1 XIF = tested non-carrier of Factor X1 |
| CIT | Citrullinemia (accumulation of ammonia and other toxics in blood in young calves) | CNC = tested carrier of Citrullinemia CNF = tested non-carrier of Citrullinemia |
| Brachyspina | Brachyspina (causes abortion and stillborn, shortened spinal cord, long legs and abnormal organs) | BYC = tested carrier of Brachyspina BYF = tested non-carrier of Brachyspina |
| CD (direct test) | Cholesterol Deficiency , calves have reduced or no cholesterol resulting in death at a young age | CDF = tested non-carrier / free of cholesterol deficiency CDC = tested carrier of cholesterol deficiency (heterozygous) CDS = tested true carrier of cholesterol deficiency (homozygous) |
POLLED STATUS
| Polled Status | Gene | Gene & Expression Code |
| Polled (Current Indirect Test) | Indirect Test | POS= tested true polled (homozygous PP) POC = tested carrier of polled (heterozygous Pp) POF= tested free of polled |
RED ALLELES
| Coat Colour | Gene | Gene & Expression Code |
| RED | Red gene (MCR1) | RDC = carrier of red gene RDF = tested non-carrier of red gene |
| RED | Variant Red gene | VRR = not tested/determined by lineage. VRS = tested true (homozygous) *Includes BKC code. VRC = tested carrier (heterozygous) VRF = tested free. |
| BLACK/RED | Black/red gene (MCR1) | BRC = carrier of black / red gene |
| BLACK | Black gene (MCR1) | BKC = carrier of black gene |
MILK PROTEINS
| Milk proteins | Gene | Gene & Expression Code |
| Beta Casein (A2) | CSN2 | A1A1 = homozygous for the A1 allele A1A2 = heterozygous for A1 AND A2 A2A2 = homozygous for A2 |
| Kappa Casein | CSN2 | KCAA = homozygous for the A allele KCAB = heterozygous for A & B KCBB = homozygous for B KCAC = heterozygous for A & C KCAE = heterozygous for A & E |
HAPLOTYPES AFFECTING EMBRYO FERTILITY
Six defective haplotypes, identified via genomic testing, are known to negatively impact fertility in the Holstein breed. These are HH1, HH2, HH3, HH4, HH5 and HH6. Similarly, the JH1 haplotype is reported the Jersey breed, as is BH2 in Brown Swiss and AH1 and AH2 in Ayrshires.
The reasons why haplotypes impact fertility are unknown, however it is thought that inheritance of the same defective haplotype from each parent results in failed conception or early embryonic death.
Animals carrying one version of a defective haplotype are coded with a C. For example, an animal carrying HH1 is coded HH1C. Animals identified as having no copies (i.e. free of the haplotype) are coded with a T, for example HH1T.
HAPLOTYPE INHERITANCE
1. If both parents are carriers of an undesirable haplotype (HH1C):
There is a 25% chance that there will be an affected offspring that would not survive to birth
Of the live offspring, one-third will be unaffected non-carriers and two-thirds will be carriers, for example: HH1C Cow (carrier = Rr) x HH1C Cow (carrier = Rr) R = Normal haplotype
r = HH1 Haplotype (containing the causative mutation)
2. If the dam is unknown, but the grandsire and bull are both unaffected carriers of an undesirable haplotype (HH1C): There is a 12.5% chance that the resulting embryo will not survive to birth
3. If the cow and the bull were carriers of different haplotypes, e.g. if the cow was HH1C and the bull was HH2C, the following resulting offspring could be expected:
- 25% non-carriers of both (HH1T and HH2T) )
- 25% carriers of one (HH1C)
- 25% carriers of the other (HH2C)
- 25% carriers of both (HH1C and HH2C)
BREEDING DECISIONS AND FERTILITY HAPLOTYPES
The best advice is to avoid using bulls which carry any of these defective haplotypes, as using such sires will reduce conception rates in the average herd. If the females being mated have also been genomic tested, then a carrier bull can be used if the female is a non-carrier. However, breeders may not want to risk producing a carrier calf, which may have a lower financial value for breeding, for example if it was an AI candidate bull.
APPLYING FOR A TEST
Genetic conditions can be identified via either genomic testing or a stand-alone test. To order a test please contact Holstein UK Membership Services on 01923 695200.